LightSwitch™ Luciferase Assay System Overview
System components:
- LightSwitch™ Promoter and 3′UTR GoClone Collections
- RenSP: an improved novel luciferase reporter gene and vector
- LightSwitch™ Luciferase Assay Reagents
The LightSwitch™ Luciferase Assay System is a unique collection of over 30,000 cloned regulatory elements from the human genome. The system includes a novel luminescent reporter gene (RenSP) and vector, an optimized companion assay reagent, and pre-cloned promoters and 3′UTRs.
LightSwitch™ Promoter and 3′UTR GoClone Collections
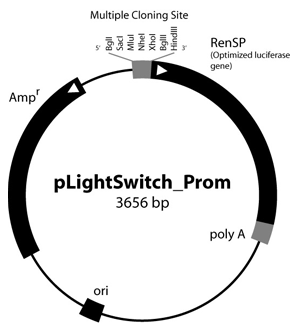
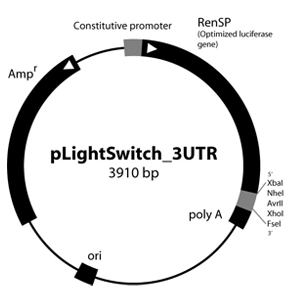
GoClone® reporter vectors are transfection-ready vectors, so no cloning or DNA preparation is required. Our GoClone® Collections include more than 18,000 human promoters and 12,000 3′UTRs cloned into LightSwitch reporter vectors (Figure 1).
Search for GoClones
Understand the basics of promoter reporter experiments.
Understand the basics of 3′UTR reporter experiments.
Figure 1: LightSwitch™ Vector Maps
Functionally Validated Pathway Sets
SwitchGear also offers functionally validated sets of promoter and 3′UTR reporter constructs for measuring a variety of biological responses. Figure 2 shows experimental results from one of our validated sets — the glucocorticoid receptor target promoters in response to three anti-inflammatory drugs. Figure 3 shows the functional knockdown of four human 3′UTRs in response to the presence of the microRNA mir-122.
 Figure 2: Glucocorticoid pathway profiling panel of promoter reporters
Figure 2: Glucocorticoid pathway profiling panel of promoter reporters
The promoter constructs shown in the heatmap represent the SwitchGear Glucocorticoid Receptor Profiling Plate. The heatmap summarizes the untreated activity of each promoter in the left column. On the right side of the figure, the log2 ratios of treated/untreated activity are shown for 3 different compounds according to the color scale shown at the base of the heatmap; “DEX”= HT1080 fibrosarcoma cells treated with 100 nM dexamethasone for 4 hours; “PRED”= HT1080 fibrosarcoma cells treated with 1 uM prednisone for 4 hours; “CORT”= HT1080 fibrosarcoma cells treated with 1 uM cortisone for 4 hours.
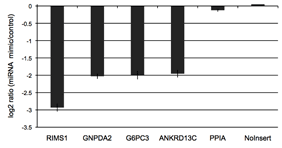
Figure 3: Validated 3′UTR targets of miR-122
The putative 3′UTR targets of miR-122 were initially predicted by the miRanda algorithm. We co-transfected putative miR-122 targets with either 20nM of miR-122 mimic or a non-targeting control miRNA in K562 cells. We selected the four strongest responders from this experiment along with two non-responding controls.
RenSP: a novel luciferase reporter gene and vector
Marine bioluminescense has evolved independently many times, so luciferases that target the same substrate (coelenterazine) bear little resemblance to one another (8). Renilla reniformis is a sea pansy that responds to mechanical stimulation by generating a blue-green bioluminescense (8). The small size of its native luciferase gene and protein (936bp and 36kD) and its lack of dependence on ATP provide a distinct advantage over larger ATP-dependent luciferases like those from fireflies (~1.6kb and 62kD) (9-12).
Why Renilla?
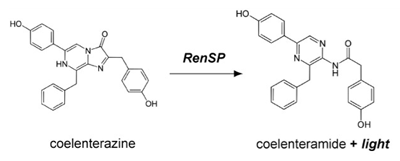
Marine luciferases have become popular alternatives to firefly luciferase as a genetic reporter based on assay simplicity, high sensitivity, and a broad linear range of signal that provides greater sensitivity over firefly luciferases (6,7). The Renilla luciferase protein catalyzes oxidation of its coelenterazine substrate in the reaction shown below to produce light at 480 nm, easily read by standard luminometers (13).
Optimizing Renilla for our novel reporter assays – increased enzymatic activity and brightness
 SwitchGear has created an optimized Renilla luminescent reporter gene, called RenSP, by increasing its overall enzymatic activity (light output) and adding a protein destabilization domain to decrease the half-life of the RenSP protein.
SwitchGear has created an optimized Renilla luminescent reporter gene, called RenSP, by increasing its overall enzymatic activity (light output) and adding a protein destabilization domain to decrease the half-life of the RenSP protein.
Starting with a base sequence of the native Renilla gene, we functionally screened thousands of synthetic gene sequence variants that included a variety of predicted improvements. We also removed transcription factor binding sites from the gene sequence that might confound expression measurements. As a result, we created the RenSP luciferase that is 100% brighter than other humanized versions of Renilla luciferase (Figure 4) (7).
Figure 4: Absolute signal of RenSP is significantly brighter than hRlucP 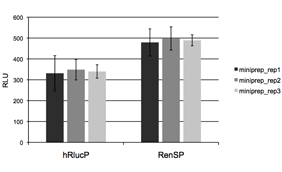 To determine the relative brightness of RenSP compared to hRlucP, another humanized form of the Renilla luciferase, the RenSP and hRlucP genes were cloned into separate vectors each containing the human RPL10 promoter. Three independent plasmid purifications were conducted for each vector, and 50ng of each plasmid was transfected with FuGENE HD in triplicate in human HT1080 cells in 96-well format. After 24 hours of incubation, 100uL of LightSwitch Reagent was added to each well and incubated for 30 minutes before being read for2 seconds on an LmaxII-384 luminometer.These results show that RenSP is significantly brighter than hRlucP.
To determine the relative brightness of RenSP compared to hRlucP, another humanized form of the Renilla luciferase, the RenSP and hRlucP genes were cloned into separate vectors each containing the human RPL10 promoter. Three independent plasmid purifications were conducted for each vector, and 50ng of each plasmid was transfected with FuGENE HD in triplicate in human HT1080 cells in 96-well format. After 24 hours of incubation, 100uL of LightSwitch Reagent was added to each well and incubated for 30 minutes before being read for2 seconds on an LmaxII-384 luminometer.These results show that RenSP is significantly brighter than hRlucP.
The RenSP gene has been fused to a protein destabilization domain (the PEST sequence from mouse Ornithine Decarboxylase (mODC) shown to increase rates of protein turnover (14-17)) to reduce protein accumulation that interferes with the ability to detect expression changes. The RenSP-PEST fusion protein therefore combines the benefits of increased signal with a short half-life reporter to provide a sensitive measure of the induction or repression of reporter gene activity. Figure 5 highlights an example in which signal knock-down of a UTR reporter after addition of a miRNA is more easily detected when the target UTR is fused to a PEST-containing reporter gene.
Figure 5: RenSP with PEST increases the knock-down of 3′UTR targets in the presence of mir-122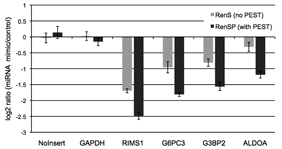 The knockdown of 3′UTR-luciferase activity in the presence of mir-122 was measured by co-transfecting a 3′UTR GoClone reporter with a synthetic miRNA. DharmaFECT DUO was used to transfect HT1080 cells in triplicate in 96-well format with 100ng of 3′UTR reporter and 20nM of mimic or non-targeting control miRNA. After 24 hours of incubation, 100ul of LightSwitch Assay reagent was added to each well, plates were incubated at room temperature for 30 minutes and read on a LmaxII-384 luminometer. The log2 ratio of the average mimic signal divided by the average signal from the non-targeting control was calculated and shows that the RenSP luciferase with PEST gives a significantly stronger knockdown than RenS without PEST.
The knockdown of 3′UTR-luciferase activity in the presence of mir-122 was measured by co-transfecting a 3′UTR GoClone reporter with a synthetic miRNA. DharmaFECT DUO was used to transfect HT1080 cells in triplicate in 96-well format with 100ng of 3′UTR reporter and 20nM of mimic or non-targeting control miRNA. After 24 hours of incubation, 100ul of LightSwitch Assay reagent was added to each well, plates were incubated at room temperature for 30 minutes and read on a LmaxII-384 luminometer. The log2 ratio of the average mimic signal divided by the average signal from the non-targeting control was calculated and shows that the RenSP luciferase with PEST gives a significantly stronger knockdown than RenS without PEST.
LightSwitch™ Luciferase Assay Reagent
We formulated a novel proprietary LightSwitch™ Assay Reagent in parallel with our gene improvements to create a fully optimized reporter system. Historically, coelenterazine-based luciferase assays faced challenges associated with high background readings. The combination of the LightSwitch Assay Reagents and the RenSP reporter gene overcomes these issues to provide:
- Bright and stable signal over many orders of magnitude with low background levels (Figure 6).
- Convenience: Add reagent directly to cultured cells (in FBS-containing or FBS-free media) in a single step.
- Flexibility: Use for a small number of samples or thousands of assays in a large screen.
- Optimized results: RenSP is optimized for use with LightSwitch Assay Reagents, providing robust results for all phases of your experimental workflow.
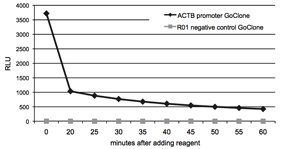 Figure 6: Signal stability of LightSwitch Assay Reagents
Figure 6: Signal stability of LightSwitch Assay Reagents
To determine the signal stability of LightSwitch Reagents, HT1080 cells were transfected with FuGENE HD in 96-well format with the ACTB promoter GoClone and the R01 negative control GoClone in separate wells. After incubating for 24 hours, 100uL of LightSwitch reagent was added to each well and luminescence was recorded for 2 seconds at 5 minute intervals during a 60 minute time-course on an LmaxII-384 luminometer. The light output stabilizes after the initial luminescent flash common to all coelenterazine-based luciferases.
References
1. Trinklein, N., Force Aldred, S., Saldanha, A., and Myers, R. 2003. Identification and functional analysis of human transcriptional promoters. Genome Res. 13:308-312.
2. Trinklein, N., Force Aldred, S., Hartman, S., Schroeder, D., Otillar, R., and Myers, R. 2004. An abundance of bidirectional promoters in the human genome. Genome Res. 14:62-66.
3. Cooper, S., Trinklein, N., Anton, E., Nguyen, L., and Myers, R. 2006. Comprehensive analysis of transcriptional promoter structure and function in 1% of the human genome. Genome Res. 16:1-10.
4. Trinklein, N., Karaoz, U., Halees, A., Force Aldred, S., Collins, P., Zheng, D., Zhang, Z., Gerstein, M., Snyder, M., Myers, R., and Weng, Z. 2007. Integrated analysis of experimental data sets reveals many novel promoters in 1% of the human genome. Genome Res. 17:720-731.
5. Collins, P., Kobayashi, Y., Nguyen, L., Trinklein, N., and Myers, R. 2007. The ets-related transcription factor GABP directs bidirectional transcription. PLoS Genet. 3:e208.
6. Lorenz, W., Cormier, M., O’Kane, D., Hua, D., Escher, A., and Szalay, A. 1996. Expression of the Renilla reniformis luciferase gene in mammalian cells. J. Biolumin. Chemilumin. 11:31-37.
7. Zhuang, Y., Butler, B., Hawkins, E., Paguio, A., Orr, L., Wood, M., and Wood, K. 2001. A new age of enlightenment. Promega Notes. 79:6-11.
8. Haddock, S., Moline, M., and Case, J. 2010. Bioluminescence in the sea. Ann. Rev. Mar. Sci. 2:293-343.
9. Matthews, J., Hori, K., and Cormier, M. 1977. Purification and properties of Renilla reniformis luciferase. Biochemistry. 16:85-91.
10. De Wet, J., Wood, K., Helsinki, D., and DeLuca, M. 1985. Cloning of firefly luciferase cDNA and the expression of active luciferase in Escherichia coli. Proc. Natl. Acad. Sci. 82:7870-7873.
11. De Wet, J., Wood, K., DeLuca, M., Helsinki, D., and Subramani, S. 1987. Firefly luciferase gene: structure and expression in mammalian cells. Mol. Cell. Biol. 7:725-737.
12. Lorenz, W., McCann, R., Longiaru, M., and Cormier, M. 1991. Isolation and expression of a cDNA encoding Renilla reniformis luciferase. Proc. Nat. Acad. Sci. 88:4438-4442.
13. Hori, K., Wampler, J., Matthews, J., and Cormier, M. 1973. Identification of the product excited states during the chemiluminescent and bioluminescent oxidation of Renilla (sea pansy) luciferin and certain of its analogs. Biochemistry. 12:4463-4468.
14. Rogers, S., Wells, R., and Rechsteiner, M. 1986. Amino acid sequences common to rapidly degraded proteins: the PEST hypothesis. Science. 234:364-368.
15. Loetscher, P., Pratt, G., and Rechsteiner, M. 1991. The C terminus of mouse ornithine decarboxylase confers rapid degradation on dihydrofolate reductase. Support for the PEST hypothesis. J. Biol. Chem. 266:11213-11220.
16. Ghoda, L., Sidney, D., Macrae, M., and Coffino, P. 1992. Structural elements of ornithine decarboxylase required for intracellular degradation and polyamine-dependent regulation. Mol. Cell. Biol. 12:2178-2185.
pGL4 Luciferase Reporter Vectors Technical Manual. #TM259. Promega Corporation.